UC Santa Barbara researchers recently studied the biophysics of the regulation and aggregation of Tau, a protein critical in regulating several intracellular functions and believed to play a role in the development of Alzheimer’s disease.
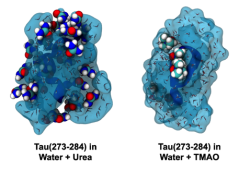
Solvent and cosolvent surfaces around Tau in the presence of urea (left) and TMAO (right). Urea binds more tightly relative to TMAO. Image courtesy of Zachary Levine
The human nervous system produces a unique set of proteins called intrinsically disordered proteins (IDPs), which can shapeshift to alter their function. These IDPs, unlike most other proteins in the body, lack a defined three-dimensional structure and are capable of transforming into a specific configuration depending on their chemical environment. Tau, a building block of specialized cells, is one of these IDPs that stabilizes the microtubules that serve as the skeleton of the neuron.
Proteins are biological macromolecules made up of amino acids, whose order and frequency determines the final folded shape of the protein. Globular proteins, which are proteins that adopt a defined three-dimensional structure on their own, have different sequences from intrinsically disordered proteins. IDPs depend on binding with a partner molecule before adopting a particular structure.
Tau’s morphological flexibility gives the cell certain advantages, but the protein can also aggregate into clumps known as neurofibrillary tangles within the neuron, causing Alzheimer’s disease and other dementias.
Joan-Emma Shea, a professor in the Chemistry and Biochemistry department and principle investigator in the study (along with MCDB professor Stuart Feinstein), said that understanding Tau’s aggregation and folding behavior may provide insights into how to treat neurological diseases.
“Cytotoxic amyloid fibrils, constructed from misfolded amyloid proteins like Tau, tend to accumulate the most on cellular membranes, but no one really understands how or why,” Shea said. “We want to understand what roles membranes and other organic surfaces play in the formation of amyloid fibril and to better understand what factors prevent or encourage their formation.”
Zachary Levine, a UCSB postdoctoral researcher in the Department of Chemistry and Biochemistry and Department of Physics, said that IDPs are largely hydrophilic, or “water-loving,” which allows them to adopt extended conformations.
“IDPs tend to have few hydrophobic, or water-fearing, amino acids,” Levine said. “Electrical interactions are therefore very important for intrinsically disordered proteins.”
The researchers used two different osmolytes, urea and trimethylamine N-oxide (TMAO), to alter the conformations of proteins. They investigated the competition between one type of electrical interaction called hydrogen bonds, which occur between a hydrogen bound to an electronegative atom and another highly electronegative species, and salt-bridges, which occur between charged amino acids. For normally-folding proteins with a fixed three-dimensional structure, the hydrophobic parts of the amino acid sequence are buried in the core of the protein to allow for a stable equilibrium between these hydrogen bonds and salt-bridges.
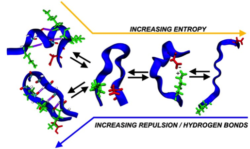
The more intramolecular hydrogen bonds present, the fewer possible configurations the protein can adopt at the cost of lower entropy. Image courtesy of Zachary Levine
According to Shea, Tau’s unique ability to shift between binding conformations made it a desirable target for osmolyte studies.
“Intrinsically disordered proteins, however, are continually frustrated, frequently shifting back-and-forth between compact states that are rich in hydrogen bonds, to intermediate states which are rich in salt-bridges,” Shea said. “More extreme situations can arise when Tau lacks both of these interactions, where it is free to sprawl out into more extended geometries.”
Urea naturally occurs in humans and is used as a protein denaturant, and trimethylamine N-oxide (TMAO) is in saltwater organisms that stabilizes compact proteins structures.
Shea said that the group found that urea halts aggregation, whereas TMAO encourages Tau to form stable, compact conformations.
“We observed that the protein could be modulated between compact and extended geometries, which was significant because Tau has a tendency to clump together inside neurons and disrupt normal cellular function,” Shea said. “However, by encouraging Tau to take on more extended structures in the presence of urea, these clumps did not form, whereas clumps could be created more easily in the presence of TMAO.”
Extended forms of Tau can link together to form a long sheet of aggregates that lead to neurofibrillary tangles. When the system is exposed to urea, urea binds to the extended structures and interferes with the sticking together of the Tau proteins. When TMAO is introduced instead, the osmolyte encourages the proteins to compact into helical structures, a precursor to large Tau aggregates.
“Adding urea breaks hydrogen bonds, which makes compact conformations unfavorable,” Levine said. “Adding TMAO, on the other hand, redistributes water around the protein exterior, which makes intramolecular hydrogen bonding more favorable.”
The researchers ran computer simulations to analyze how osmolytes change Tau’s shape and its ability to aggregate. Their results suggested that the structure and aggregation of the IDP was related to how tightly bound the protein is to other Tau proteins.
“The chemical structure of urea is quite similar to the backbone of Tau,” Levine said. “Urea mimics the protein structure and binds to the surface, stopping other pieces of Tau from binding to one another because the binding sites are already occupied by urea.”
These IDP aggregation studies may contribute to further understandings of protein-related diseases. Their findings were published in the Proceedings of the National Academy of Sciences.















