The word “stochastic” is not a commonly used word. The word only recently entered the vernacular of chemists and biologists when studying the organisms living in our gut, also called our microbiome. So what does it mean? “[A stochastic process is] like rolling a die … [it means] it’s random,” said Eric Jones, a postdoctoral researcher at Simon Fraser University in British Columbia, Canada, who received her doctoral degree in physics at UC Santa Barbara.

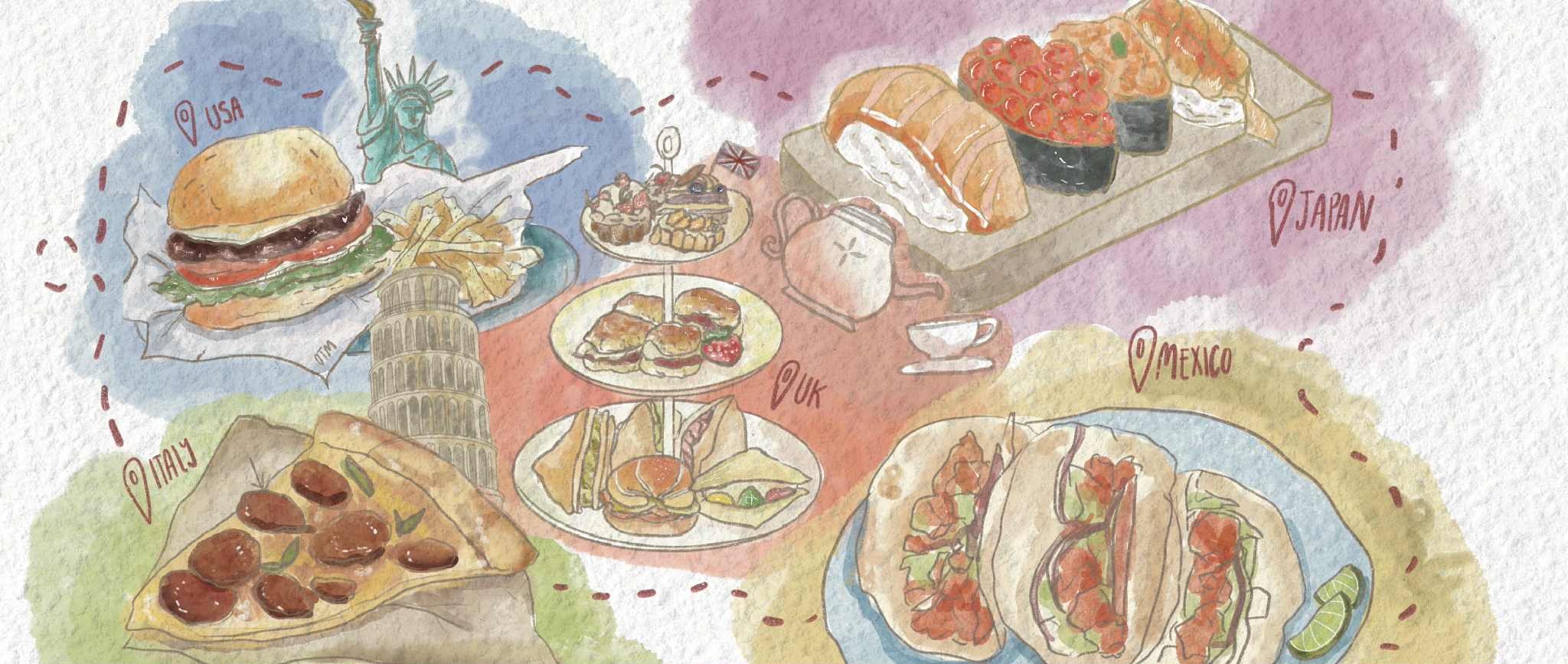
The success of one bacteria implanting into a microbiome has been tough for scientists to measure. Courtesy of National Human Genome Research Institute
The stochastic model of implantation by organisms — such as by taking up semi-permanent residence in your microbiome — is a recent development in the field.
Researchers do this by applying probabilities to the likelihood of a given species implanting in a biome.
“You could say, for some species, let’s define successful colonization only if you roll a one. Whereas for other species that are stronger colonizers, maybe they’ll colonize so long as you roll a one, two, three or four,” Jones elaborated.
And in their new paper in “Proceedings of the National Academy of Sciences,” titled “Stochastic microbiome assembly depends on context,” Jones, UCSB physics professor Jean Carlson and their colleagues discussed the introduction of the notion of context to the field. Jones was a doctoral student in the physics department with Carlson when he started to work on the published material. Instead of focusing on the individual successes of isolated microorganisms implanting in a microbiome, the report looked at how the presence of other microorganisms affects the process.
“The context part means that the weighing of the die itself is affected by which other species [are already present],” Jones explained. “[I]n effect, [nature] is rolling a die that is no longer fair.” This is the context present to a given species when it is given the opportunity to implant in a microbiome. A given species will have different odds of colonizing a microbiome if other species are present. Jones and Carlson introduced a weighted statistical method to quantify this contextual model of implantation.
According to Jones, the human microbiome is difficult to experimentally probe as there are thousands of species present in our microbiome at any given time. William Ludington’s group at the Carnegie Institution for Science and Johns Hopkins University Department of Biology instead looked at the microbiome of fruit flies that were bred to be born with no bacteria initially present in their microbiomes. Ludington is a co-author on the paper and his group contributed all the experimental data found in the report. The flies were fed a yeast compound with a known dosage of any combination of five bacteria species from the genera Lactobacillus and Acetobacter. The Ludington group meticulously fed a specific fly only one of these compounds, and each of the bacterial combinations was fed to 48 different flies. The microbiomes of the fruit flies were then sequenced to see how successful each species was at impacting the microbiomes.
With 48 data points for each of the 31 possible combinations of bacteria, Jones and Carlson were able to apply a stochastic- and context-dependent statistical model to the probabilities of a given bacteria species impacting the microbiome. They compared their new method to the success of older models of species success rates and looked at the difference between their experimental results and theoretical predictions. Each of the models was considered to have “explained” a bacterial species combination if the model predicted a success rate within two standard deviations of the experimental data. Independent models that do not rely on context predicted 20 of the 31 combinations. Jones and Carlson’s context-dependent model predicted 24 of the 31.
Underlying the urgency to this new analysis is the booming field of probiotic treatments and fecal transplants to positively alter human microbiomes. Current therapies are similar to the experiments conducted in Ludington’s lab — some work and some don’t. Not all fecal transplants are created equal; some people get categorized as “super-donors.”
“They have really good poop,” Jones said. “Really good at colonizing and grafting in other people’s guts. [M]aybe these super-donors have some bacteria that are strong colonizers like we’ve seen in our experiments.”
Two people can receive fecal transplants with the same goal — to implant a given bacteria species — and one could succeed while the other could fail.
The stochastic model is a scientific way of explaining random successes and failures. Jones and Carlson’s new context-dependent model will hopefully expand the field’s thought process on how to improve microbiome therapies. The success of one bacteria implanting into a microbiome has been tough for scientists to measure.
“When you say how successful a fecal transplant is, it’s hard to quantify,” Jones said. “[You] have a sick person’s gut made up of a thousand species of bacteria, and each of those species has some abundance. It’s a big vector.”
With a context-dependent model, interactions between different species can be quantified and applied in designing better fecal transplants. A bacterial species not initially requested or required for a fecal transplant could now be important in the success of the therapy if it aids in impacting another desired bacterium. The context-dependent model could be a paradigm shift in this field.
“That’s the hope,” Jones said. “It would be fair to say that this is not really how people thought about [fecal transplants].”