For many, the word “biodiversity” springs to mind images of equatorial rainforests and coral reefs, big expansive habitats filled to the brim with plants and animals of such a wide array of shapes, sizes, roles and lineages that one cannot even comprehend their true breadth.

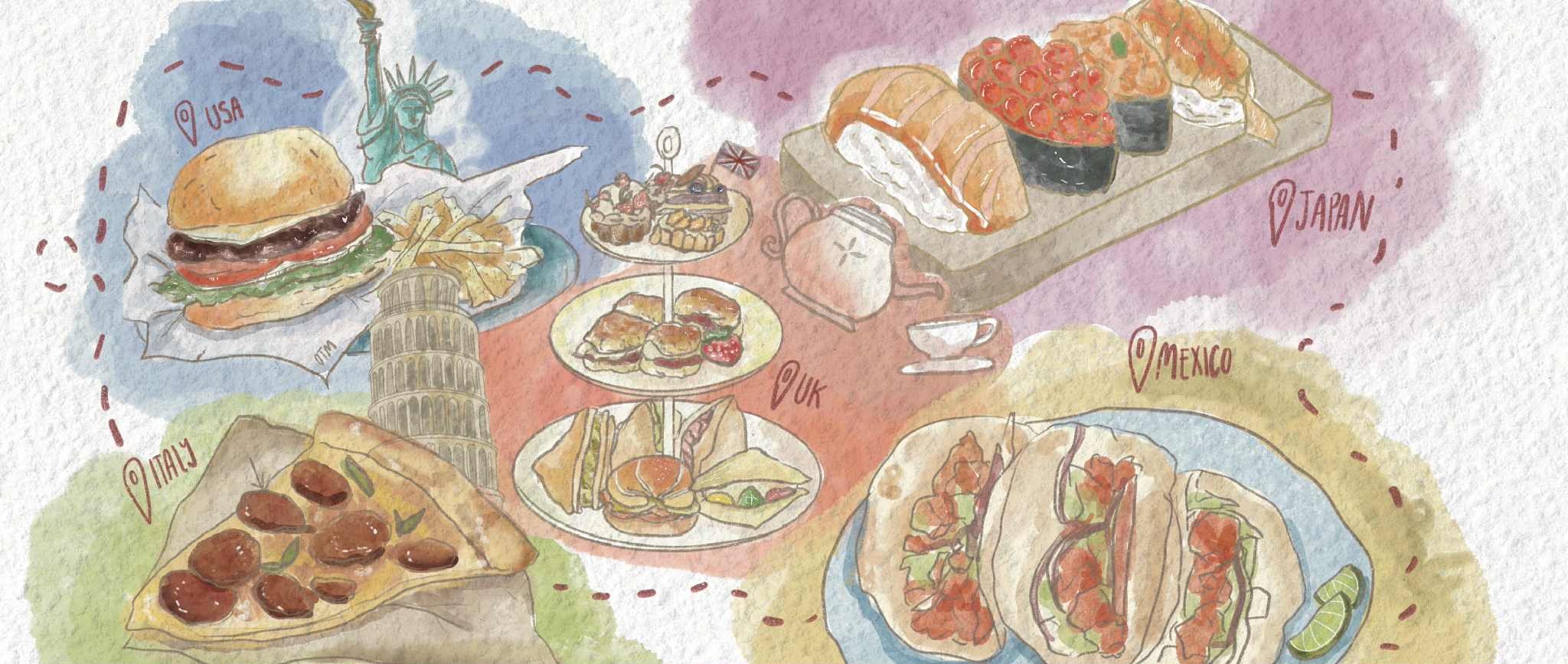
A flock of sheep. Kabsik Park / flickr
And within these habitats are presumed to be a wealth of natural products useful for their pharmaceutical, therapeutic and antimicrobial potential. These are the subjects of intense interest and study.
But biodiversity is a descriptor that fits more settings than just the megadiverse regions found in the tropics. The California floristic province, for instance, where Isla Vista is found, stretches from Baja California to Southern Oregon and is the home of some 3,400 taxa of endemic plants.
We can even go more local than that, however. Look within individual animals — the kind one can find at a petting zoo, like sheep, goats and horses — and a whole world of microbial diversity opens up. Consider microbiomes: the totality of microbiota living on and within vertebrates.
“We know that there are natural products that give us highly valuable medicines and therapeutics that are sourced from really exotic environments,” Candice Swift, a researcher at the UC Santa Barbara Department of Chemical Engineering, said.
“But something as simple as the digestive tract of a goat or a sheep is actually really complex and a really interesting environment to look at … We actually have these sources of microbially sourced chemical diversity that are right there at our fingertips.”
To raise just one example, UCSB researchers in the chemical engineering department, Michelle O’Malley and Swift, and their cross-university collaborators have been hard at work peering into the gut microbiome of large herbivores, cataloguing the genes within these which encode biosynthetic enzymes.
For the researchers’ samples, they used representative species of the thousands of species of gut fungi found in large herbivores. Four were selected, Anaeromyces robustus, Caecomyces churrovis, Neocallimastix californiae and Piromyces finnis, isolated from canary grass enrichment from sheep and goat feces in the Santa Barbara Zoo and the feces of a horse from a farm in Massachusetts.
From this, the researchers carried out a series of omics analyses at four different levels, looking at the genome, the mRNA, the proteins and the enzymes produced by the fungi and the final small molecules derived from those sources.
“So there’s a lot of precedent in the literature about natural products. People know that there are these particular enzymes that make chemically diverse and interesting small molecules that are medicinally relevant like antibiotics,” Swift said.
The researchers decided to look at the DNA of these representative species to identify the enzymatic diversity — and in turn, the chemical potential — within the fungi.
Then, the researchers looked at the active genes through the mRNA of the representative species to understand the other biomolecules which are tied to the final small molecule. Swift carried out sequencing experiments to characterize under what conditions these genes within the fungi may be expressed.
In conjunction with this, Swift’s partners with the Environmental Molecular Sciences Laboratory in the Pacific Northwest National Lab characterized the actual proteins and enzymes of these genomes to better understand what they do.
Finally, the small molecules themselves were characterized.
“It’s a good idea to study biological information at these four different levels, because the level of complexity just gets higher and higher as you go to the final small molecule. [If all] we have is this metabolic soup of small molecules, it’s a lot harder to say whether this small molecule is going to be a polypeptide versus a nonribosomal peptide, for instance,” Swift said.
Meanwhile, “at the level of DNA, it’s really easy to tell those different types of enzymes apart that make those small molecules.”
Through this analysis, the researchers found a host of genes (146 in total) which encode for biosynthetic enzymes and in turn natural products.
For Swift, such a discovery is particularly exciting because it’s a new concept in general that these animals may have anaerobic fungi which make natural products at all. According to Swift, their research is the first of its kind in scientific literature.
In fact, in general, the digestive tracts of large herbivores haven’t really been studied in the context of drug or antibiotic potential for decades, according to Swift.
“And there’s nothing to the level that we have done where we’ve basically unraveled the whole chemical repository of these organisms and shown that it’s very much under-appreciated in the context of natural products,” Swift said.